 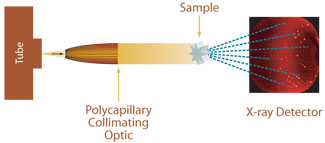
Three major technologies are available to make protein structures ‘visible’:Īs more than 90% of all protein structures deposited in the publicly accessible protein database of biological macromolecules w3 have been determined by X-ray diffraction, we will concentrate on this method. Particles that size cannot be observed even with the strongest light microscope, which has a maximum resolution of 1 micrometre (1 m = 1 thousandth of a mm). Proteins are tiny structures, measuring only a few nanometres (1 nm = 1 millionth of a mm). Image courtesy of Beat Blattmann and Patrick Sticher Proteins are too small for direct observation Workflow for protein structure determination by X-ray diffraction Nevertheless, since they are involved in fundamental biological processes, there is a great interest in better understanding their structure and function, and scientists keep trying to crystallise them. Due to their complexity, these proteins are experimentally extremely challenging, and every time the structure of a protein is determined, it is a major achievement. One of the major challenges in structural biology today is the elucidation of the structure, function and interaction of huge macromolecular complexes and membrane proteins w2. Investigating the structural details of a protein is of great importance to understand how fundamental processes of life function at a molecular level: this is the research area of structural biologists. These sites can catalyse biochemical reactions, as in the case of enzymes, or form a specific binding site, as in the case of antibodies. Only when the protein is folded, the specific amino acids of the protein are close enough to enable the formation of an active site. The function of a particular protein depends on its three-dimensional structure. Structure is function: what does the three-dimensional structure of a protein tell us? The entire protein forms a tertiary structure consisting of a variety of such structure elements. The most prominent elements are α-helices and β-sheets (see figure below), which are typically stabilised by hydrogen bonds between individual amino-acid residues. Stretches of amino acids form typical secondary structural elements. Under natural conditions, the linear chains of amino acids spontaneously fold into distinct three-dimensional structures. Proteins are folded into distinct three-dimensional structures The assembly is performed by a ribosome, which is a complex molecular machinery consisting of proteins and RNA. In cells, each protein is assembled using the information encoded in its corresponding gene. The length of the protein chain varies from a few dozen to thousands of amino acids. They consist of 20 different building blocks, called amino acids, which are arranged in a linear chain connected by covalent bonds between adjacent amino acids (see figure below). The variety is immense: more than 20 000 different proteins are known to exist in humans alone.ĭespite this variety, all proteins share an identical structural principle. antibodies) or the transport of small molecules (e.g. hormone receptors), immune responses (e.g. collagen in connective tissue), or mediate cell signalling (e.g. Other types of proteins have mechanical and structural functions (e.g. Almost every biochemical reaction requires a specific protein, called an enzyme. Proteins are the largest group of non-aqueous components in living cells. Fifty years later, however, it is still a challenge to obtain protein crystals for structural studies. 
This pioneering work was awarded the Nobel Prize in Chemistry in 1962 w1. They grew crystals from this protein and managed to determine its structure by analysing the X-ray diffraction pattern of the crystal.Ī number of myoglobins from other species had been tested before with little success, until Perutz and Kendrew obtained a useable diffraction pattern with whale myoglobin crystals. By investigating the protein’s structure, the two scientists wanted to understand the oxygen-carrying mechanism at the molecular level. In 1959, Max Perutz and John Kendrew published an article on the three-dimensional structure of whale myoglobin, which is a small protein responsible for the transport of oxygen in whale cells. Author(s): Beat Blattmann, Patrick Sticherīeat Blattmann and Patrick Sticher from the University of Zürich, Switzerland, explain the science behind protein crystallography and provide a protocol for growing your own crystals from protein – an essential method used by scientists to determine protein structures. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
